Research helps producers select cultivars and provides plant breeders tools to ensure that blackleg major gene resistance lasts
The canola industry introduced a labelling scheme for major blackleg-resistant gene groups used in canola varieties back in 2017, but until recently there was no data from Canadian fields on how effective this approach is.
Justine Cornelsen is an agronomy specialist at the Canola Council of Canada, as well as a grad student at the University of Manitoba. She’s studying the strategic deployment of blackleg-resistant gene groups in commercial canola fields on the Canadian prairies.
The study focuses on the use of resistant cultivars and the rotation of these resistant sources.
Read Also

Growing garlic by the thousands in Manitoba
Grower holds a planting party day every fall as a crowd gathers to help put 28,000 plants, and sometimes more, into theground
“Within this particular project our research objectives were to validate this concept of blackleg major gene resistant label cultivar deployment,” Cornelsen said during a presentation at the University of Saskatchewan’s annual Soils and Crops workshop that was held online this year.
Avirulence genes are genes of a pathogen that govern its specific recognition by particular plant genotypes. A plant’s recognition of the pathogen depends upon the presence of a pair of matching genes, an avirulence gene in the pathogen and a resistance gene in the plant.
The study Cornelsen presented examined blackleg incidence and severity and it was also designed to develop empirical data on how the avirulence changes within fields, to help understand how effective major resistance gene cultivar deployment can be in Canada.
“This is something that we have seen in other canola producing regions, but it has never been validated here in our environmental conditions. But we’ve seen it work well in greenhouse and growth chambers and trials here, but not in our general growing landscape,” Cornelsen said.
She said the project had a large survey component that collected information on the cultivars farmers grew and how they managed their fields in the 2018 and 2019 growing seasons.
The study’s goal was to have 10 fields from each of the Prairie provinces, but a few of the fields were lost due to dry conditions.
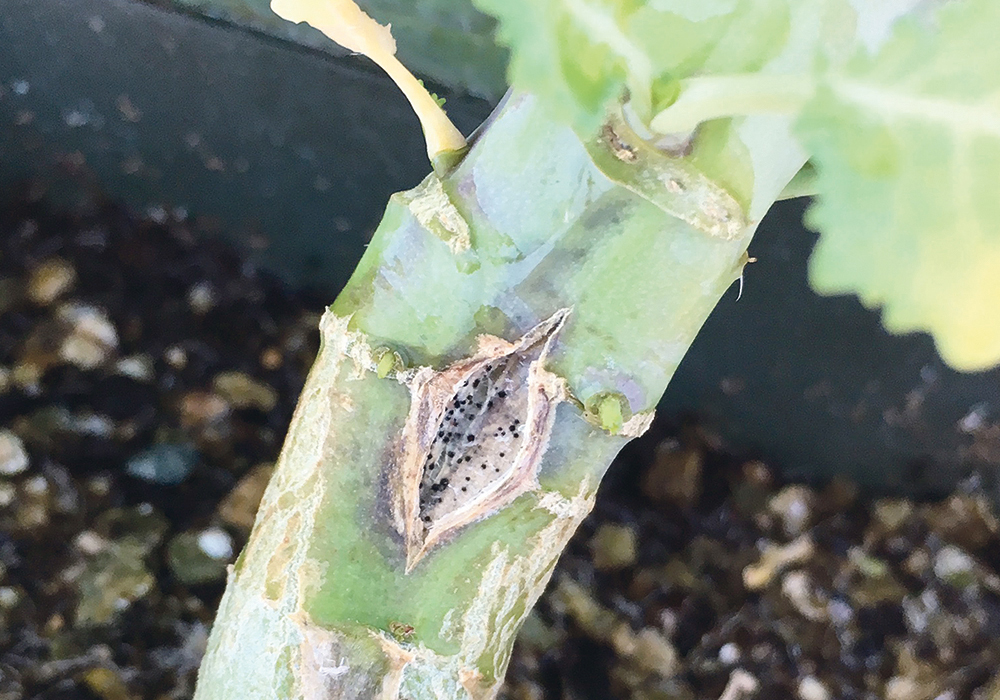
“Old residue was collected in the spring prior to canola planting and then that field of canola that was planted was then rated for disease severity and incidents, and more samples were collected so we could figure out what the avirulence profile of the blackleg or of leptosphaeria maculans was in the field,” Cornelsen said.
Leptosphaeria maculans is a fungal pathogen that is the causal agent of blackleg.
Researchers documented what the cultivars had in terms of their major resistance genes, and then ran an avirulence profile of the infected residues with PCR and also cotyledon inoculation tests.
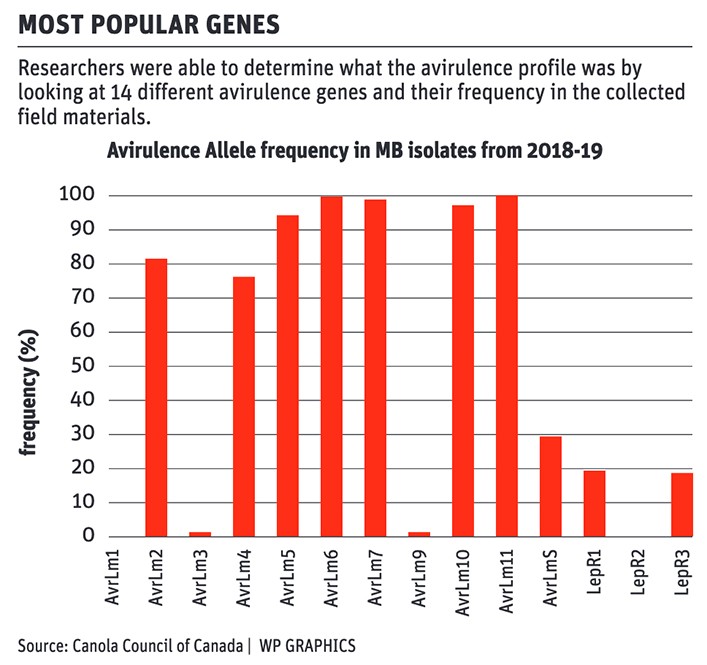
“We were able to capture and figure out and determine what the avirulence profile was and that was done looking at 14 different avirulence genes,” Cornelsen said.
Disease severity was rated on a zero to five scale, and then disease incidents and disease severity was broken down and looked at based on the major resistance gene groupings.
There were six different major resistance gene group combinations made up of four major resistance genes that were commercially available to canola producers when the study was conducted.
“What we were looking at is to try to see if there was differences between deployments. So both within disease incidence and severity there was significant differences between the resistance gene groups,” Cornelsen said.
She said it was also important to understand what the leptosphaeria maculans race profile was.
“Based on the disease incidence we can then look at the samples from the field in the springtime and see why we were getting the disease incidence and severity within the different cultivars,” Cornelsen said.
She provided an example of Saskatchewan fields surveyed in 2018 where any of the cultivars that contained major gene resistance RLM4 had significantly lower disease incidences and severity.
“When we look at the avirulence profile of leptosphaeria maculan from the spring this makes sense. There is avirulence4 present in the population and you can see it’s just over 80 percent of the isolates collected were containing that,” Cornelsen said.
She said avirulence3 wasn’t found in any of the isolates because it was suppressed by the presence of the avirulence seven, and that RLM3 hybrids had 70 percent disease incidence and higher disease severity because their resistance didn’t match up with the blackleg races present in the field.
The study found more than 35 unique blackleg races in Manitoba, however, Cornelsen said there are lots of resistance genes available that match to all of the races commonly found.
The study also looked at the allele breakdown in the blackleg population.
An allele is a variant form of a gene.
Cornelsen said avirulence4 is found in 70 percent of the samples collected in Manitoba so the RLM4 resistance gene is a good option on many fields in the province.
RLM3 is very common in available canola varieties, however it is not a common allele found on Manitoba fields so there is a low chance it will be effective.
Cornelsen said the information collected in the study will help producers select cultivars and provide plant breeders with avirulence frequencies and ensure that blackleg major gene resistance lasts, as well as help reduce trade concerns of blackleg inoculum in seed exports.
The R/MR/MS/S rating system for blackleg remains in place that rates the quantitative resistance, which is also known as minor gene resistance or background resistance of canola varieties.